Diffusion MRI allows us to trace white-matter tracts in vivo by detecting the random thermodynamic motion of water molecules in brain tissues. With this technique, it is possible to reconstruct the human connectome at a millimeter scale by modeling each brain region as a network node and white-matter pathways as network edges. Using functional MRI, we construct the functional networks by modeling the edges as the correlation between the time series of two brain regions. Both human structural and functional networks present a complex topology, such as small-world architecture, rich-club, modularity, and a cost-efficiency balance. We seek to fully describe the organization principles of both human structural and functional networks using tools from graph theory and algebraic topology.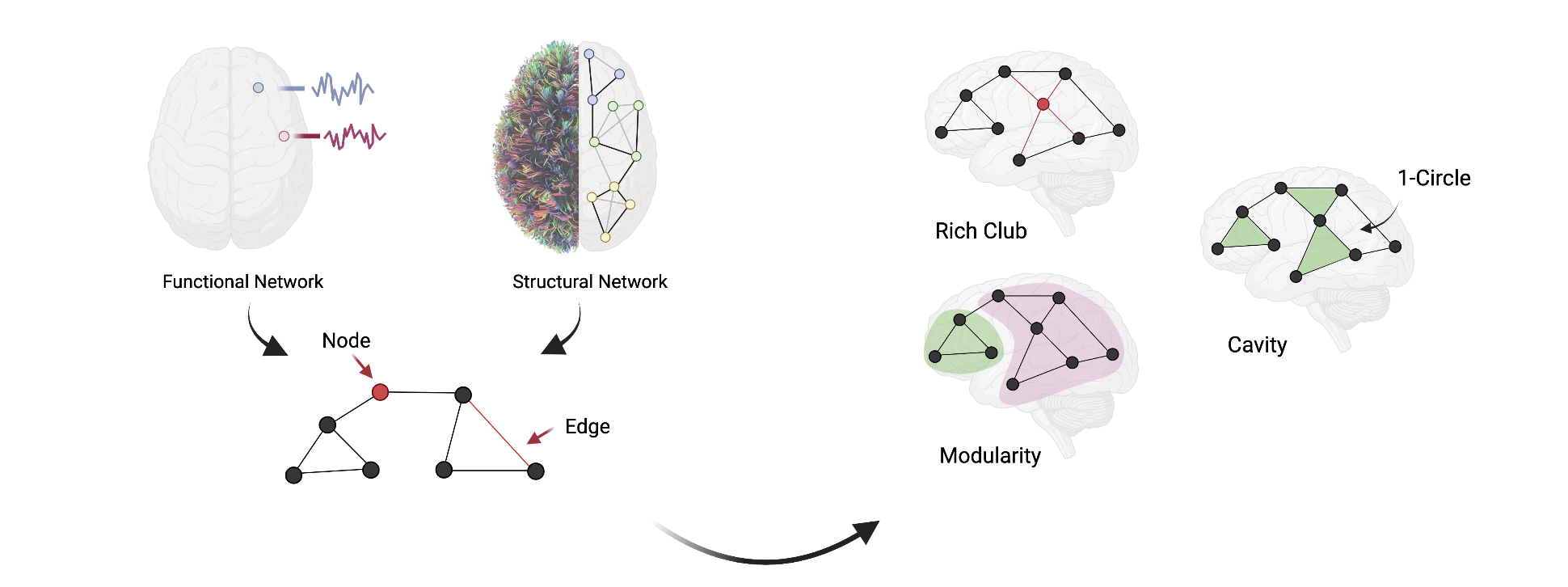 The network topology of structural connectome supports the communication dynamics between neuronal elements, which underlies all aspects of brain function and human cognitions. We are interested in building a generative model of how the functional dynamics during the execution of a cognitive task emerge from the underlying structural connectome. To link connectome and cognition, we employ computational tools from network control theory and implement experimental manipulation with both whole-brain non-invasive neuroimaging (i.e., fMRI and MEG) and local invasive recordings (i.e., iEEG).
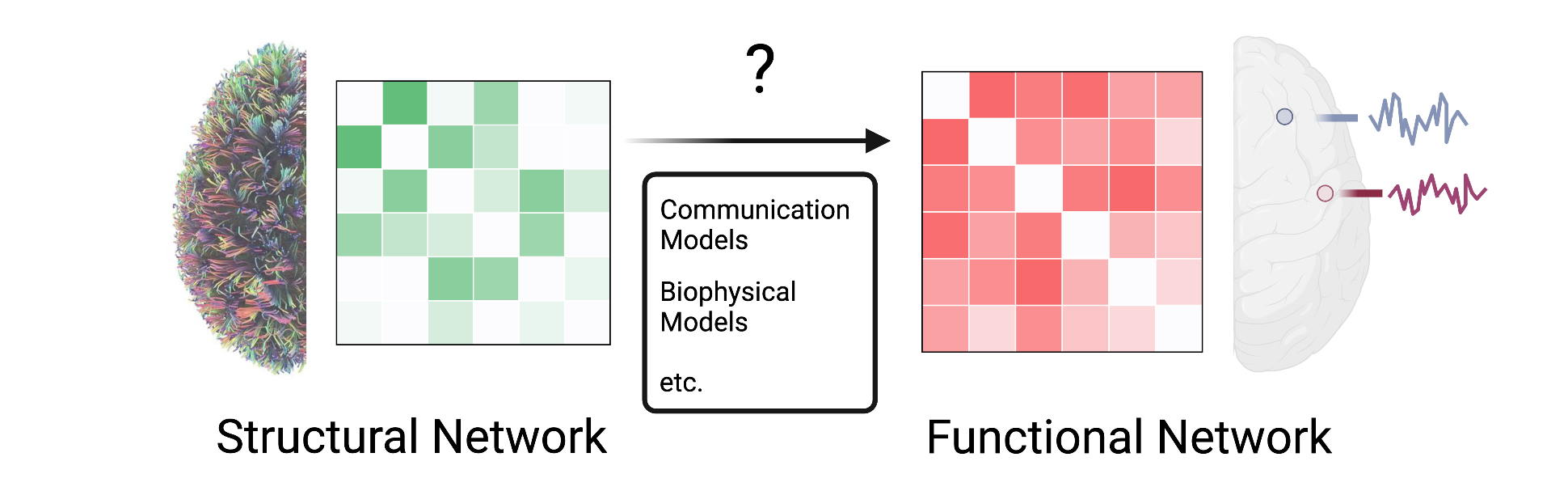
The network topology of structural connectome supports the communication dynamics between neuronal elements, which underlies all aspects of brain function and human cognitions. We are interested in building a generative model of how the functional dynamics during the execution of a cognitive task emerge from the underlying structural connectome. To link connectome and cognition, we employ computational tools from network control theory and implement experimental manipulation with both whole-brain non-invasive neuroimaging (i.e., fMRI and MEG) and local invasive recordings (i.e., iEEG).
 TOP
TOP